
Recent advances in deep learning have dramatically enhanced the accuracy and efficiency of medical image registration. Yet, such algorithms often struggle to generalize beyond the image types they see at training.
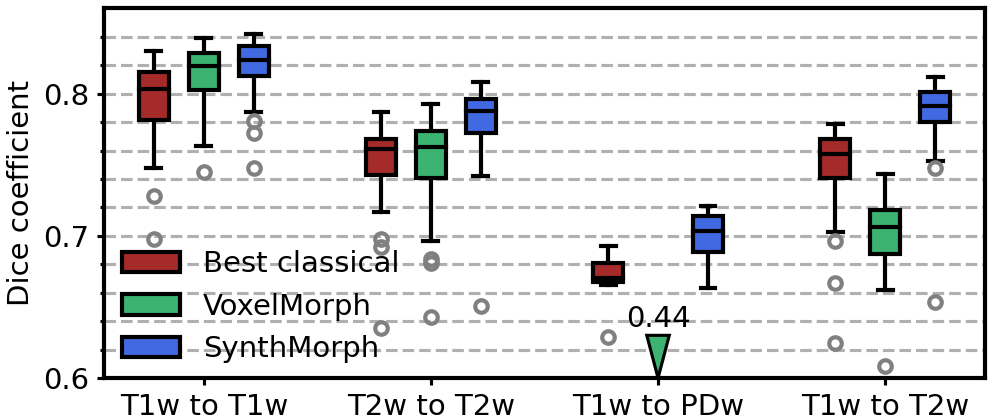
The SynthMorph learning strategy for acquisition-agnostic registration addresses this challenge by exposing networks to widely variable synthetic data, which leads to unprecedented robustness across a landscape of real image types and greatly alleviates the need for retraining.
SynthMorph is also an easy-to-use registration tool.
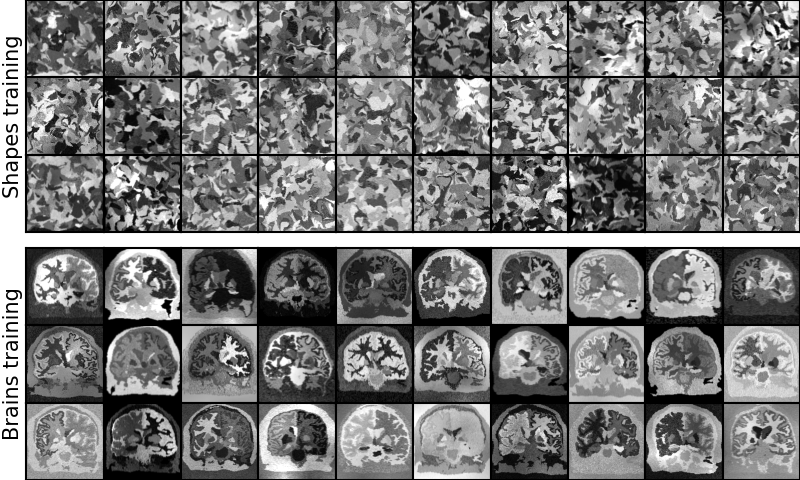
SynthMorph is also a tool leveraging the synthesis strategy for rigid, affine, and deformable registration of brain MRI without preprocessing such as skull-stripping. The tool registers image volumes of any size, resolution, orientation, and contrast. The moving and the fixed image geometries can differ, and you can change the warp regularity at test time.
We maintain a standalone container version on the Docker Hub, with a script for easy setup and use supporting the following container tools: Docker, Podman, Apptainer, or Singularity.
In addition,
FreeSurfer
ships the command-line tool mri_synthmorph. For the latest version,
download a build
of the FreeSurfer development branch.

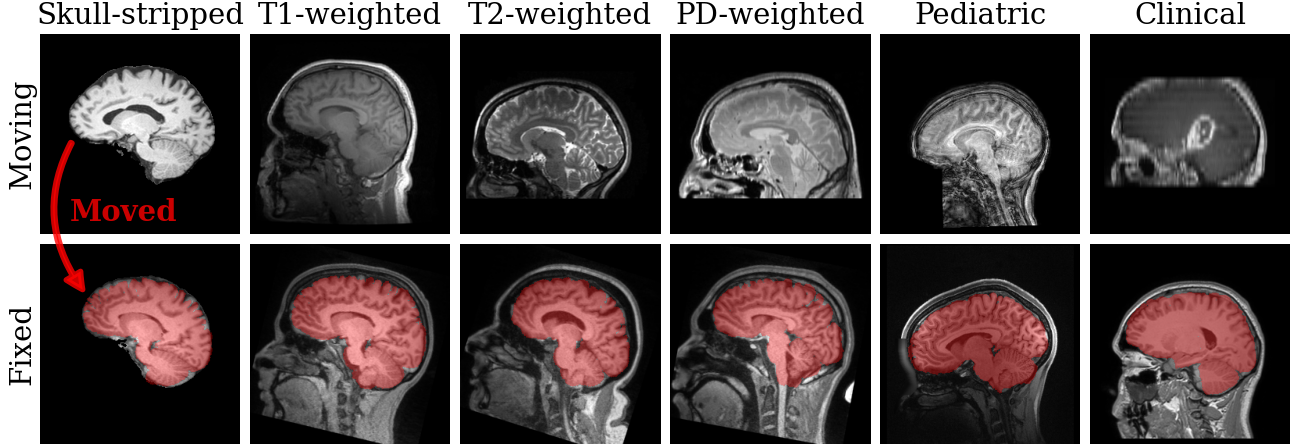
Register a moving to a fixed image using the default joint
affine-deformable transformation model and save the result as
moved.nii:
mri_synthmorph -o moved.nii moving.nii fixed.nii
Run the deformable registration at 25% regularization strength,
saving the joint displacement field as trans.nii:
mri_synthmorph register -r 0.25 -t trans.nii moving.nii fixed.nii
Estimate and save an affine transform trans.lta
in FreeSurfer LTA format:
mri_synthmorph register -m affine -t trans.lta moving.nii fixed.nii
Apply an existing transform to an image:
mri_synthmorph apply trans.lta moving.nii.gz out.nii
Apply an existing transform to a label map:
mri_synthmorph apply -m nearest trans.nii labels.nii out.mgz
Print detailed instructions and information on transforms:
mri_synthmorph -h
For those interested in training registration models, we maintain Google Colab notebooks showcasing the SynthMorph learning strategy:
Users who wish to build custom environments can find SynthMorph scripts in FreeSurfer, access the Python requirements for the latest container, and browse source code on GitHub. The weight files are available under MIT or CC BY 4.0 license—choose either:
Several papers describe SynthMorph registration techniques, where * denotes equal contribution. If you use SynthMorph, please cite the relevant papers below (BibTex).
Joint registration and toolbox:
Affine and rigid registration:
Deformable registration:
Synthesis-driven training and domain randomization:
Start a discussion, open an issue on GitHub, or reach out to the FreeSurfer list.
We thank Danielle F. Pace for help with computing surface distances. SynthMorph benefited from computational hardware generously provided by the Massachusetts Life Sciences Center.